The Neurogenomics and Informatics (NGI) Center (https://neurogenomics.wustl.edu/) is requesting proposals for a new pilot grant program. The vision of the NGI is to transform the field of human neurogenomics by going beyond the analysis of genomic DNA to explore other omics layers, including epigenomics, transcriptomics, proteomics, and metabolomics. Molecular phenotyping of human samples is instrumental […]
Author: cruchagac
MS4A4A is the major regulator of TREM2 levels
Drs. Harari, Benitez, Karch and Cruchaga leveraged biospecimens obtained from large and well-characterized human cohorts to identify a novel protective gene for Alzheimer disease, MS4A4A, that is also the major regulator of TREM2. This study provides a strong evidence of a biological link between TREM2 and MS4A4A in microglia in the context of AD. However, […]
Drs. Cruchaga and Karch receive new funding to advance personalized medicine in Alzheimer Disease
Drs. Cruchaga and Karch are some of the Washington University investigators that received funding from Centene to perform molecular phenotyping of Alzheimer’s cases to identify novel molecular biomarkers and the identification of novel therapeutically targets. Specifically we plan to develop a personalized medicine approach to understand the effects of Alzheimer’s disease risk genes by combining […]
Novel protective variants for Alzheimer’s Disease risk identified
A study led Dr. Kauwe lab and in collaboration with Drs. Karch and Cruchaga identified rare variants in RAB10 that protects against Alzheimer’s Disease risk. In this study Dr. Karch lab performed the functional studies and Dr. Cruchaga lab contributed genetic data. A video explaining the findings can be found below:
Using Genetics to predict risk for AD at individual level (Links to an external site)
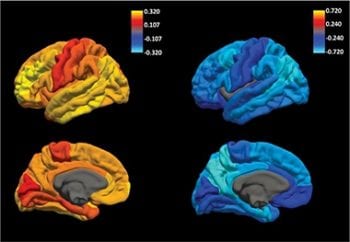
Dr. Cruchaga discuss about the use of polygenic risk scores to predict individual level risk for Alzheimer and provide this data “direct-to-consumer”.
Mendelian Neurodegenerative mutations and 23&me (Links to an external site)
In collaboration with Alzforum Dr. Cruchaga analyses the impact of direct-to-consumer genetic testing.
Our work presented at the AAIC2017 was highlighted by Alzforum (Links to an external site)
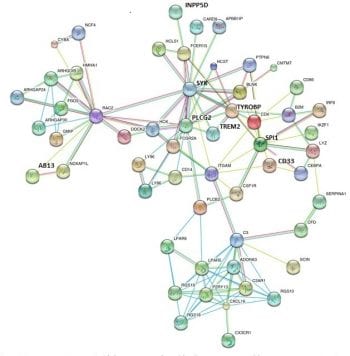
Geneticists are inventing new ways to hunt AD variants that went undetected in GWAS and might shed light on AD pathogenesis. Many labs are searching for polymorphisms tied to specific quantitative traits. As Yuetiva Deming from Carlos Cruchaga’s group at Washington University, St. Louis, pointed out, GWAS identify risk variants but say nothing about how that risk manifests
MS4A Alzheimer’s Risk Gene Linked to TREM2 Signaling (Links to an external site)
Leveraging IPS-derived neurons and brain tissue to identify novel genes implicated on Frontotemporal Dementia (Links to an external site)
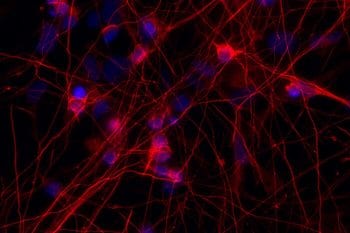
R406W causes a specific form of frontotemporal dementia. We found that GABAergic dysfunction in postmortem tissue from people with sporadic FTD or progressive supranuclear palsy, but not in Alzheimer’s disease. Also highlighted in Alzforum: https://www.alzforum.org/news/research-news/stem-cell-model-nails-link-between-tauopathy-and-gabaergic-dysfunction
Analysis of TREM2 structure (Links to an external site)
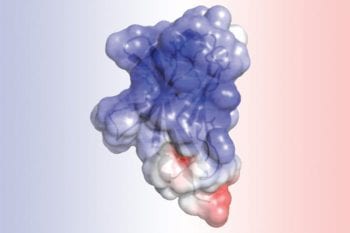
Great collaboration with Dr. Brett and Colona lab
Cruchaga lab got more than $7 million from NIH to study Alzheimer’s genetics (Links to an external site)
The studies are funded by two grants totaling $7 million from the National Institute on Aging of the National Institutes of Health (NIH) and led by Carlos Cruchaga, an Professor of psychiatry and of neurology at the School of Medicine
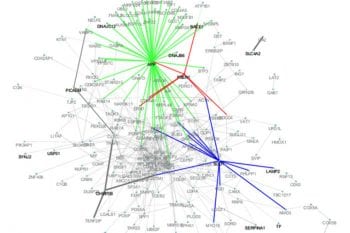
Harari and Cruchaga lab develop a new digital deconvolution for brain RNA-seq data (Links to an external site)
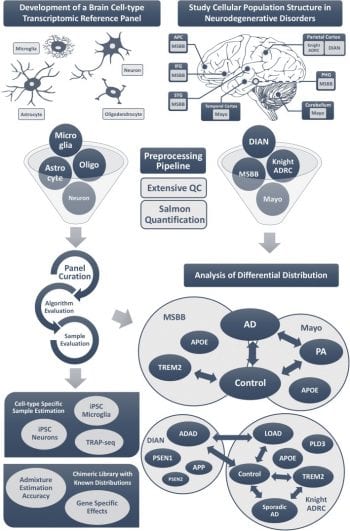
Our computer method determined the proportions of each cell type — neurons, astrocytes, oligodendrocytes, microglial cells — and found that specific gene variants were linked to different proportions of these cell types.
The NeuroGenomics and Informatics leaders receive funding from the Chan Zuckerberg Initiative (Links to an external site)

Drs. Harari, Cruchaga and Karch received funded from the Chan Zuckerberg Initiative to understand the relationship between MS4A4A and TREM2